
Hi! My name is Yepeng Du, I am a PhD student from the Taipale group at the Faculty of Medicine, University of Helsinki.
During my masters, I was using SELEX (Systematic Evolution of Ligands by EXponential enrichment) technique to select single-stranded DNA aptamer, characterizing screened aptamer and developing aptasensor (aptamer-based biosensor) against erythromycin for real-time detection of polluted water sources because chemistry based methods, exemplified by HPLC, are generally time-consuming even though they can be greatly sensitive and accurate.
Now, I am working on my PhD and I’m studying to decrease cancer cells’ reliance on Myc oncogene for the purpose of controlling cancer growth.
What role does Myc oncogene play in cancer development?
To put it as simple as possible, to control cancer growth is nearly equal to controlling expression of the necessary Myc responsive genes.
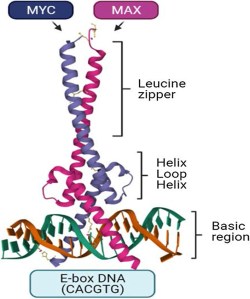
The genesis of most human cancers (>70%) is contributed from aberrant expression level of transcription factor Myc that may be able to regulate at most 15% of the whole genome via dimerizing with other transcription factors and binding to E-box (CACGTG) within respective controlled gene, whereby various cellular processes are directly regulated, including transcriptional regulation, biosynthesis, cell cycle control and signaling transduction.
While some Myc responsive genes are essential to cellular fitness, others are not, because the latter, with decreased expression, do not have a negative impact on cell growth.

In addition to this, it has not been reported yet that an existing specific drug is able to antagonize Myc protein, due to its ‘undruggable’ features that are granted by its lack of enzymatic pocket available to bind with conventional small molecules.
Therefore, an anti-Myc antibody is not capable of accessing the nucleus and neutralizing Myc protein through nuclear pore. Considering these points, the reasonable measure might be to make other transcription factor to replace Myc protein.
How to make it come true?
CRISPR/Cas9 mediated gene editing assay was utilized to replace E-box (promoter of genes controlled by Myc) contained in important Myc controlled genes of cancer cells to other host transcription factor’s binding motif, by which we could see what transcription factor can transactivate the edited genes.
Seyed Esmaeil Ahmadi, Samira Rahimi, Bahman Zarandi, Rouzbeh Chegeni, and Majid Safa.
J Hematol Oncol. 2021; 14: 135. Creative Commons license 4.0. No changes to the image were made.
Why reduce the dependence on Myc, rather than to mute Myc oncogene?
Given that such an important characteristic of Myc is its role in cellular fitness, we cannot simply “delete” Myc gene or mute its expression, otherwise normal cell activity will be negatively influenced as well.
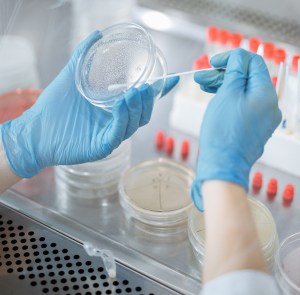
In brief, gene edited cells were cultured for a few days, followed by being collected to prepare an Illumina library for checking if they grew or not. This would prove if the newly introduced motif worked or not.
Gene edited cells were also cultivated for preparing ChIP-qPCR library that could tell if the edited genes were truly transactivated or transrepressed by the new motif.
What is the future work?
To further reduce the role of Myc in malignant cells, a dTag system will be employed with CRISPR/Cas9 gene editing assay to conduct this research.

Want to learn more about the Finnish Center of Excellence in Tumor Genetics?
Keep up to date with the latest tweets, videos and blog posts for a peek into the everyday life in cancer research by following us on Instagram (@tumorgenetics) and Twitter (@CoEinTG). You can also subscribe to our YouTube channel!