By Sinja Taavitsainen and Matti Annala
Cancer affects the lives of tens of millions of people every year. The high incidence of cancer has motivated research into more effective treatments to increase patient lifespan and quality of life. Cancers have traditionally been treated with surgery and drugs that indiscriminately eliminate rapidly dividing cells in the patient’s body. However, clinicians now also have access to drugs that eliminate cancer cells based on specific molecular traits that distinguish them from normal cells. These targeted therapies can have higher efficacy and better tolerability than older drugs, but are only effective in patients whose cancer cells harbor a specific molecular weakness. Their use therefore requires knowledge of the molecular mechanisms driving an individual patient’s cancer.
Historically, studying cancer cell genomes during the later stages of disease has been challenging because this requires tissue biopsies to be obtained from metastases in the patient’s body. These biopsies are invasive to the patient, carry a risk of complications, and are particularly difficult to obtain in cancers that primarily metastasize to bones (such as prostate cancer). In recent years, “liquid biopsies” based on circulating tumor DNA have emerged as an alternative to tissue biopsies.
What is circulating tumor DNA?
All humans carry short circulating DNA fragments, known as cell-free DNA, in their blood. These DNA fragments originate from damaged or dying cells in the body. In cancer patients, a significant fraction of cell-free DNA in the blood can originate from tumor cells. These tumor-derived DNA fragments are referred to as circulating tumor DNA. By capturing and reading the circulating tumor DNA present in a patient’s blood, scientists can characterize the DNA mutations that drive a patient’s cancer.
The amount of circulating tumor DNA in a patient’s bloodstream depends on the number of cancer cells in their body (among other factors). It is often quantified as the percentage of circulating tumor DNA, relative to total cell-free DNA in the sample. This percentage is referred to as the circulating tumor DNA fraction (ctDNA fraction, ctDNA%). If the circulating tumor DNA fraction is very low, it may be impossible to detect cancer driving mutations in the blood sample, and a traditional tissue biopsy may be required instead. In healthy people, 10 mL of blood typically carries a quantity of cell-free DNA equivalent to the diploid genomes of 6000 cells. Patients with metastatic cancer can sometimes harbor 100 times higher concentrations of cell free DNA in their blood, so that over 90% of the cell-free DNA in their blood originates from cancer cells.
What does the use of liquid biopsies mean for a cancer patient?
Liquid biopsies promise to provide genomic information about a patient’s disease with minimal risk and invasiveness. A liquid biopsy requires only 5 – 10 mL of blood to be taken from the patient, enabling multiple samples to be collected during the course of their disease. In prostate cancer, there are now drugs known as PARP inhibitors coming to use that are highly effective for the 20% of patients whose cancer cells carry specific mutations disrupting DNA repair, but have no benefit for the remaining 80% of patients. Rising circulating tumor DNA levels in a liquid biopsy can also be used to detect cancer recurrence after treatment.
What does the use of liquid biopsies look like in practice?
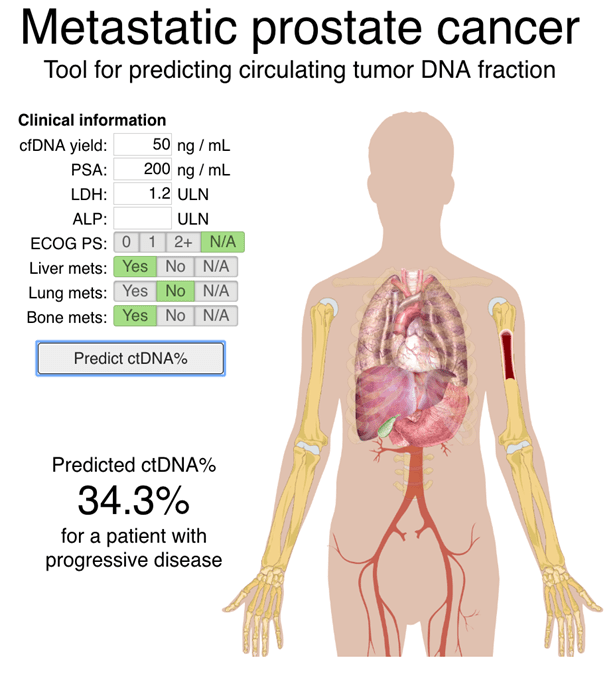
During the past several years, the use of liquid biopsies in both the research and clinical setting has become increasingly common. This growth is fueled by continued improvement in our ability to extract useful genomic information from these samples. In the Computational Biology group at Tampere University, we are developing computational methods for more sensitive and accurate analysis of circulating tumor DNA biopsies. Together with our collaborators in Canada, we recently published the first study comparing time-matched cfDNA and tissue biopsies in a cohort of 45 metastatic prostate cancer patients, proving that cell-free DNA biopsies provide a more practical alternative to tissue biopsies in many patients. In related work, we recently developed a model for predicting the circulating tumor DNA fraction in metastatic prostate cancer patients based on routinely measured clinical variables such as prostate specific antigen (PSA) levels. We have made this tools available as a website.
In collaboration with clinicians and researchers from Canada, United States, and Finland, we analyze cell-free DNA samples collected from patients participating in randomized clinical trials to understand the molecular mechanisms that drive resistance against cancer drugs. In one recent clinical trial, we analyzed cell-free DNA samples from 202 patients that were randomized to receive one of two hormone therapies, abiraterone or enzalutamide. Liquid biopsies were collected from the patients at three timepoints: before taking either drug, after progressing on the first drug, and after progressing on the second drug. We analyzed the circulating tumor DNA to determine how mutations in the cancer genome relate to drug response. This was the first large prostate cancer trial in the world where cell-free DNA biopsies were obtained from all patients. We are now finalizing a second manuscript about this clinical trial, where we are now focusing on how the cancer cells change when they acquire resistance against these widely used drugs. We are also simultaneously working on another clinical trial comparing cabazitaxel against hormonal therapies in metastatic prostate cancer, with cell-free DNA biopsies again collected from all patients.


What are some future directions for liquid biopsies and ctDNA analysis?
Although circulating tumor DNA-based liquid biopsies are already a great tool for research and guiding treatment, the cost and time required to sequence and analyze each sample remains a crucial factor. While sequencing and analysis currently takes 1 – 5 days, the analysis of circulating tumor DNA in the future is likely to be done in real-time. With the development of portable sequencing instruments, it may become feasible to analyze patient blood samples within clinics, allowing doctors to make decisions about patient treatment within hours or minutes of sample collection.
Finally, one of the most important endeavors in oncology is the early diagnosis of cancer before metastatic spread. The biggest challenge here is that small localized cancers shed only a tiny amount of circulating tumor DNA into the bloodstream, so that high sensitivity is paramount. Furthermore, in addition to diagnosing the presence of cancer in a patient’s body, it is often necessary to also diagnose its location for localized treatment. Since many cancer types harbor characteristic mutations, and modern sequencing technology is capable of detecting individual mutant DNA fragments in a blood sample, early diagnosis through circulating tumor DNA detection is already becoming a reality.
Another fascinating aspect of cell-free DNA biopsies is that they capture genomic information about multiple cancer cell populations in a patient’s body. This is different from tissue biopsies, which only capture DNA from a single anatomic site. Combined with the ease of obtaining multiple cell-free DNA timepoints across time, this enables us to study how different cancer cell populations expand, shrink, and evolve in a patient’s body throughout treatment.
Who is involved in this work?
Our liquid biopsy projects are collaborations between the Computational Biology group at Tampere University, the Wyatt lab at Vancouver Prostate Centre (led by Dr Alexander Wyatt), and the Clinical Trials Unit at the British Columbia Cancer Agency (led by Dr Kim Chi).
