By: Riku Katainen
How did we end up in genotoxin research?
Our Tumor Genomics Group focuses mainly on acquired and inherited genetic/epigenetic changes behind tumor and cancer development in humans, so how did we end up in genotoxin research? Back in the Fall of 2019, we were contacted by Thomas F. Meyer from the Max Planck Institute for Infection Biology. They had been studying the DNA of colibactin-exposed cells and wanted us to validate their novel finding with our colorectal cancer (CRC) data. Well, why were they interested in CRC data specifically? Since it was known that colibactin would cause double-strand breaks to DNA, Meyer’s group wanted to find out if there were preferred DNA sequences these breaks would occur in.
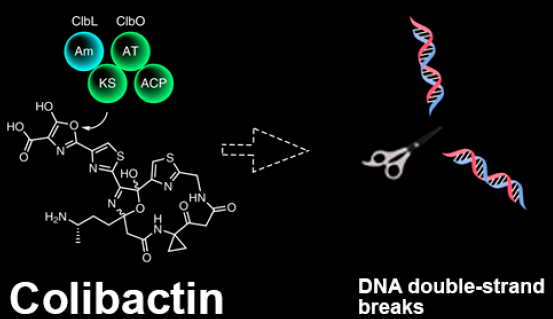
What is colibactin? Genotoxins are substances that can cause mutations to our DNA. In this research, we studied colibactin, which is a genotoxin produced by a certain strain of E. coli bacteria. E. coli bacteria normally live in the intestines of people and animals and thus makes our gut the first place, where we would expect to find cells affected by colibactin. Colibactin has been shown to induce double-strand breaks to DNA.
Indeed, there were specific short DNA sequences, motifs, which were over-represented in their DNA breakpoint analyses. The next step was to use publicly available mutation data from different cancer types to determine whether these motifs were mutated in certain cancers. They noticed elevated numbers of mutations at these motifs in a subset of the public CRC data set. Thus, our extensive and high-quality CRC sample data would be ideal for the validation of the found signature, so, they contacted and introduced us to this novel colibactin finding.
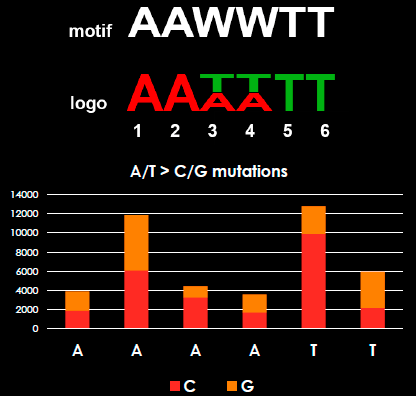
What are mutational signatures? Mutational signatures are defined by the detection of certain mutations at certain sequence contexts or motifs. For instance, tobacco smoke causes an excessive amount of C -> A substitutions and these mutations occur predominantly at ACG sequence triplets or contexts (ACG -> AAG). In this study, we show the mutational signature of A/T -> C/G substitutions, which occur at AAWWTT sequence context, where ‘W’ is ‘A’ or ‘T’ base.
Was the mutation signal true?
We promised to investigate if we see the signature in our samples, so we got a more detailed description of the found mutational signature. They had detected mutation accumulation in CRC samples at A/T (adenine/thymine) rich, six bases long sequences (hexanucleotides) e.g., AAATTT and AAAATT. At first, we were skeptical about the correctness of the finding due to the repeated nature of the motif – these kinds of sequences are generally prone to mutations and sequencing errors, which could be the cause behind the signal. Also, around 15% of CRCs are so-called microsatellite unstable, which harbor an extensive amount of mutations at similar repeated sequences. So, this finding could not be true… or could it?
Our analysis efforts refined the finding and revealed a clinical correlation
We were, fortunately, able to validate the finding – these mutations were not sequencing errors or repeat region induced, as we first feared. We developed new features to our variant analysis software, BasePlayer, and refined the motif analyses (see our earlier blog post about the software). We were able to map all mutations in our CRC data to exact positions relative to “colibactin motifs” present in the human genome. The mapping revealed, that A/T -> C/G mutations accumulated at the 2nd and 5th positions in the hexanucleotide sequences. Sublimely, these positions were exactly the same that Meyer’s lab measured in colibactin-exposed cells, which strengthened our confidence of this mutation signal to be of colibactin origin.

Next, we used these specific mutations as a variable in our regression model and detected colibactin exposure (AAWWTT mutation count divided by all genomic mutations) to be significantly more prominent at the distal parts of the colon.
Colibactin signature is more prominent in the distal colon. Figure: Riku Katainen
What’s next?
The measurements of us and others have indicated that these colibactin-induced mutations can start to accumulate already in the very early stages of life. However, the contribution of these mutations to cancer development, if any, remains to be shown. The future goals are to determine the exact mutational mechanism and to find possible pathogenic effects of colibactin. This study was published in Nature Medicine in the summer of 2020. You can read the full manuscript here.
