By Riku Katainen & Alison Ollikainen
As sequencing technology continues to advance, the amount of data that can accumulate is exponentially increasing. This means that datasets can become huge, and therefore handling and managing that data can be a daunting task, especially for those without large amounts of computing capacity and skill.
Riku Katainen, a PhD student and bioinformatician in the Tumor Genomics research group at the University of Helsinki, noticed that there was this battle, and endeavored to find a way he could make the integration, navigation and visualization of this data easier and more versatile. Not finding a software that was up to the task he decided he’d build it himself. Lovingly coined after his musical skills, his large-scale discovery tool, “BasePlayer”, was born, and even better, he made it free for everyone.
Riku tells us more…
What is BasePlayer?
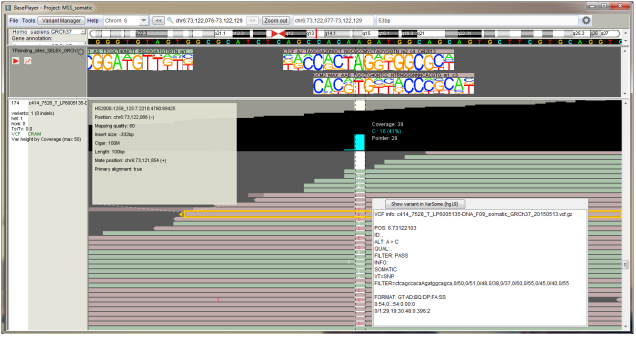
BasePlayer is a user friendly analysis software for modern disease genetics. A common example of its use is in the detection of a mutation or a gene behind the disease which is running in a studied family. Other common cases would be the detection of causative somatic mutations in a specific cancer or tumor type. Somatic mutations accumulate in all of our cells during our lifetime and after a series of unfortunate events, some of these mutations can possibly cause e.g. a tumor.
Why did you design it? Who is it for?
I designed the software, since there was no such software available that could handle large-scale mutation analysis in multiple samples visually and in a user friendly fashion. There are a multitude of analysis tools for specific tasks available, which, however, often do not have a user interface and/or require a background in computer science or bioinformatics. BasePlayer combines the features of various analysis tools and algorithms and it is designed for a general audience regardless of bioinformatics training.
What did you want to achieve by making BasePlayer
At the end of 2009 I had a vision how these kind of analyses should be done in future, so I wanted to materialize that, and due to my persistent nature and background in computer science, I was able to! 🙂
How does it connect to the rest of cancer research, how can it be used?
Modern cancer genetic research relies heavily on next-generation sequencing (NGS) data. NGS enables researchers to study the majority of an individual’s DNA information at once. Before the BasePlayer analysis, the NGS data has to be aligned to the human reference genome. The alignment enables the detection of “all” mutations in the sample’s DNA. These pre-processing steps are commonly performed by the NGS data provider. So basically, you can freely download the BasePlayer from our website, install it to your computer and open the NGS files (e.g. VCF and BAM) for analysis.
Why would you like to promote BasePlayer?
I have seen a lot of anxiety among the researchers in analysing and interpreting colossal amounts of NGS data. I would like to see a change in that, and also wish for BasePlayer and its versatility to be applied outside of our own research circles.
The BasePlayer was recently published in Nature Protocols – see the manuscript here, see full text here and the YouTube teaser here.

One thought on ““If you build it, they will come” – Innovative Tool for Mutation Analysis”
Comments are closed.